Dear blog, my source of reflection,
One thing that has occurred to me is that, in order to keep moving ahead in life, you need to routinely ask yourself why you are doing the things you do. For example, I’ve loved science since I don’t even know when, but I never really ask myself why I want to grind my nose in research or go to med school (if I even still want to). Let’s face it. I wasn’t born with an undying love for what I do. If the high school freshman I once was had seen my bioinformatics research projects that I’m currently doing, I would have gone running for the hills in any direction other than science.
I’m majoring in physics while doing research in bioinformatics with a premed track, and I’m also taking courses in philosophy, computer science, and biology. Confused? Me too. The reason I need to reflect upon these academic focuses is that I need to remember my purposes to identify where and how to move on in the next part of life.
Have I lost my focus?
When I was a freshman in high school, I fell in love with physics and mathematics. In my physics and geometry classes, the idea of putting forth a mathematical proof to describe the universe through deductive reasoning blew my mind. I began to consider the possibility of being an engineer or scientist when I grew up. I started to fantasize about how I could solve all the world’s problems through mathematics. Come sophomore year, and chemistry was my new weapon of choice. I saw this hierarchy of fundamental knowledge starting with math, then physics, and, finally chemistry. (I currently don’t believe that the universe is entirely applied math and everything can be explained through physics, but that’s beside the point). At that time in high school, I appreciated all things scientific, except for one key area in science: biology.
I was always confused out of my mind why math, physics, and chemistry were integrated so well with one another while biology was sitting at its own table with its own fantasies about the universe. I scratched my head in my introductory biology class wondering “Why on earth are we explaining this complex mechanism of photosynthesis through these arbitrarily drawn squares and circles? Does it really take a genius to sit there and say that one molecule attacks the other simply because you said so? Why are we wasting time in memorization and drawing pictures rather than reasoning and evaluating? Why are we sitting here and categorizing all the things in the world rather than actually using the categories for scientific inquiry? Where is the calculus?” Anyhow, in hindsight, I was definitely being too harsh and idealistic, like I usually am. Biology courses are not taught this way because biology is exclusive to math and physics, but, rather, because they are taught from a different angle. Albeit, I never enjoyed learning biology this way, but, nowadays, I understand that not everything has to be taught from the same scope as physics and chemistry are.
After my senior year of high school, I did a summer research program at a small, local college. The research program let me choose what I wanted to do a project about. Being the young Einstein-wannabe, I wanted to study physics or mathematics. Unfortunately, the college was only strong in biological and chemical research. But, being desperate to start a research project in anything scientifically-related for the sake of science itself, I began a project in which I studied the numbers of shrimp consumed by Hydra vulgaris. The research professor showed me the world of biostatistics, in which we use math and numbers to study biology.
This blew my mind. (Math? Statistics? And numbers? In biology? Maybe I could appreciate biology after all.) Somewhere here or there, I heard about the field of biophysics. This sounded like an amazingly perfect field for me to study because I could finally understand the elusively cryptic field of biology in something that I enjoyed.
When it was time for college, I started on a premed route (for reasons I’ll explain in the next paragraph or two. I don’t know how I’m structuring this blog post yet). I chose to major in physics just because. I also started to lurk around the university departmental websites for opportunities to join research labs. I started looking through physics labs, but, seeing as how I only had experience in biological research and statistics, I ended up joining a bioinformatics lab. This was also another check off my list of “Things I want to learn” because I’ve always wanted to learn how computer programming. During my second semester of college, I started having another affair. One night, as I was trying to fall asleep on a cold, January night, I woke up around midnight with one thought running through my mind:
Philosophy.
It made perfect sense. My love for science was not fueled by anything else other than the love of coming to solutions from problems. I chose physics because it was the most fundamental of the sciences and, frankly, the only one that really stood back and questioned the way we reason and look at the world, rather than the fields of chemistry and biology which were “structured” in their own way. I whipped out my laptop and signed up for the introductory logic course for the second semester. When I told the philosophy undergraduate advisor that I wanted to double-major in physics and philosophy, he was very pleased and excited by the reasons that I gave him. I could tell that I was unique in the way I looked at the world.
Needless to say, that introductory logic course was like a wake-up call from a thousand-year sleep.
But, if you’re paying attention, you probably remember that I’m a physics major, right? And I’m doing research in bioinformatics? Well, truth be told, I am doing physics research, too, but it’s VERY VERY slow. In fact, I haven’t accomplished anything in my physics lab during my freshman year. My physics research professor has been way too unresponsive with me, but, hopefully I can contact him at the beginning of next semester again to start getting the ball rolling. In the meantime, I found a look at this website from the IU School of Informatics and Computer Science. It seemed to be everything I’ve ever wanted to study, all in one place. I’m in the process of contacting the research professors who work for it in order to find a research opportunity that can perhaps replace my slow-moving, not-quite-dead-yet physics lab that I’m working in. Working in two research labs simultaneously shouldn’t be too big of an issue. Especially since I’m a theoretician.
Real talk. I’m a premed out of family pressure. Throughout high school, I never really found a legitimate reason for becoming a doctor, and, whenever I’d protest to my parents that I didn’t want to be a premed, they would tell me it was the best path that one could take, therefore, I should take it. They told me that going to med school and becoming a doctor was one of the highest honors that one could achieve. While studying science might get you a job for a few years, everyone would respect you as a doctor for a lifetime. But, ever since I can remember, I’ve always been idealistic. Too idealistic. In fact, only about a week ago I realized how cynically ideological my motives were that they weren’t concrete enough for you to weigh a piece of paper on it. I wasn’t obsessed by “achieving a high status in society” or “making a lot of money.” I wanted to obsess over math and science to achieve a creative bliss that transcended anything that this world could offer me. But, and maybe my parents always knew best, it’s better to have a realistic, sustainable goal in mind for the rest of your life. I never put much thought into the actual career path of being a doctor, nor into the routines and activities that doctors normally do. Among the myriad of activities that doctors do on a day-to-day basis, the only things that truly interested me from the beginning were surgery and research. Today, as I write this, I’ve found other incredibly fascinating ways to study medicine, including through informatics/computer science, policymaking, and ethical discussion. Physics, computer science, biology, and now philosophy? How on earth does medicine fit into this?
Being a pre-med and going off to study medicine has always been a love-hate relationship for me. There were times during my first semester of college when I seriously considered abandoning the track of being a doctor altogether after I realized the frustratingly structured path which denied any liberal pursuit of education. “Psychology? Why would I have to take a course in psychology?” I remember one of my close friends told me “You know, you don’t HAVE to be premed. Doctors barely do anything involving physics and mathematics anyway.” That really struck a chord with me. I couldn’t simultaneously worship the liberal studies of philosophy and physics while staying on a more conservative approach to becoming a doctor. Was it possible for me to love learning for the sake of learning and still tell myself that I wanted to be a doctor? So…why do I want to go to med school? Am I just too scared to tell my parents I don’t want to be a doctor?
I’m still looking for a reason.
I’ve come to reconcile the two worlds of research and medicine by telling people I want to do an MD-PhD program, but, even then, I need to develop a burning passion for why I want to be a doctor. And not something vague, generic, or uncultivated like “I want to help people” or “I want to find a cure for cancer.” I also can’t read the lives of other people and try to follow their footsteps. There has to be something that I do that gives me an undying love for medicine and health care like no one else has. I mean, look at this guy! He studied physics, philosophy, and mathematics during his undergraduate years and then went on to make amazing contributions to history, medicine, and so many different social sciences! Or what about this fellow? A physicist-philosopher? My dream come true! He has also written about medicine and medical ethics. Splendid.
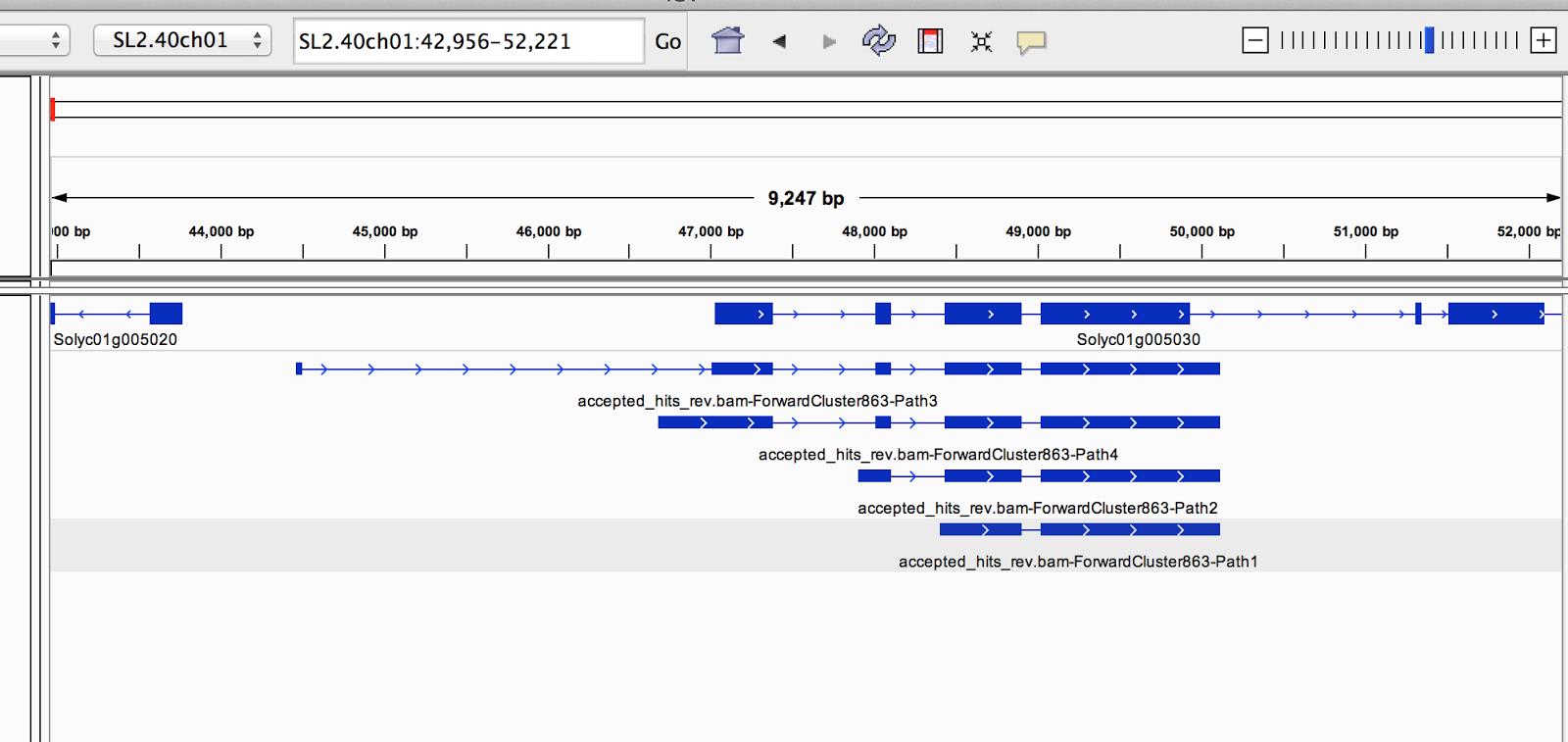
This quest is not futile. I’m sitting in my dorm room at Cornell University in the middle of my bioinformatics internship. I’m studying how the non-coding RNA regions of tomatoes do things. That’s related to health and well-being, right? It’s a good start. But I need something deeper. Something that incorporates all my interests. As far as ethics is concerned (as it relates to my love for philosophy), I am interested in medical ethics, bioethics, and scientific ethics. I can check that off the list now. Physics?…I’ll come back to that later. Hopefully. Someday. Computer science and biology? Well, those are because I became obsessed with bioinformatics.
Maybe that’s the only reason I need to justify what I study. Because I fell in love with them. Maybe I don’t need some ultimate goal of becoming the president of the country or finding the cure for cancer. I mean, I got accepted into an internship at Cornell. I think I’m doing things right, right? I just need to keep checking in on myself.
Here I go off into the world of academics, waltzing with the devilishly charming demon of scientific knowledge through physics, computer science, biology, and everything in between. Where will I end up? Let the coffee pour and spin the roulette wheel of fortune before the day’s over. We’ll see what I can learn tomorrow.
People ask me what I’m studying. Sometimes I say physics. Sometimes I say medicine. Sometimes I say computer science. And sometimes I say none of the above. Where am I going in my life? Looking back, everything has only been a serendipitous walk through a very, very confusing maze of passion after passion.